
Welcome to the
companion website of Lagarde,
Uszczynska-Ratajczak et al., Nature Communications 2016.
Long noncoding RNAs
(lncRNAs) constitute a large, yet mostly uncharacterized fraction
of the mammalian transcriptome. Such characterization requires a
comprehensive, high-quality annotation of their gene structure and
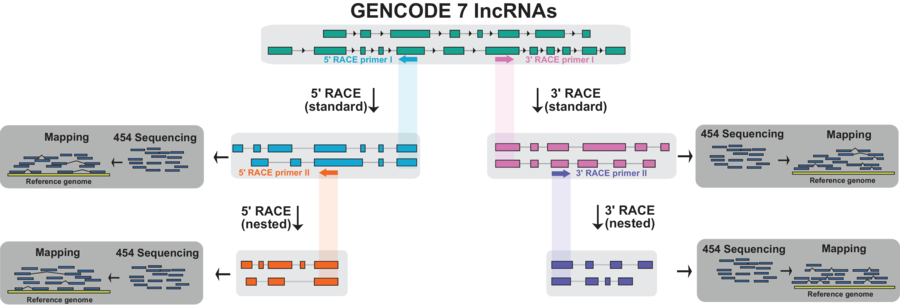
boundaries, which is currently lacking. Here, we describe
RACE-Seq, an experimental workflow designed to address this based
on RACE (Rapid Amplification of cDNA ends) and long-read RNA
sequencing. We apply RACE-Seq to 398 human lncRNA genes in seven
tissues, leading to the discovery of 2,556 on-target, novel
transcripts. 60% of the targeted loci are extended in either 5' or
3', often reaching genomic hallmarks of gene boundaries. Analysis
of the novel transcripts suggests that lncRNAs are as long, have
as many exons and undergo as much alternative splicing as
protein-coding genes, contrary to current assumptions. Overall, we
show that RACE-Seq is an effective tool to annotate an organism’s
deep transcriptome, and compares favorably to other targeted
sequencing techniques.
Detailed information about GENCODE v7 targets as well as the RACE primers used in this study can be found in the Supplementary Data of the article.
All FASTQ files generated in this experiment are downloadable from the European Nucleotide Archive (accession: ERP012249).
Track
Hubs allow convenient, in-context visualization of custom
sets of tracks in genome browsers. We have registered a Track
Hub Registry entry, with links loading the RACE-Seq track
hub directly into the UCSC Genome Browser and Ensembl.
The RACE-Seq track hub is based on genome assembly GRCh37
(a.k.a. hg19) and includes the following RACE-Seq tracks/datasets:
For each assayed
tissue, we provide a list of detected RACE-Seq transcript
models: Brain,
Heart,
Kidney,
Liver,
Lung,
Spleen,
Testis.
Collapsed, non-redundant sets of TSSs (in BED format) are linked below. "Raw" TSSs were clustered using the "bedtools merge -nms -n -s -d 50" command.
RACE-Seq: All, novel, CAGE-supported
CaptureSeq: All, novel, CAGE-supported
Questions, requests and comments about this study should be addressed to: